Combo of 3 antibiotics can kill deadly staph infections
Three antibiotics that, individually, are not effective against a drug-resistant staph infection can kill the deadly pathogen when combined as a trio, according to researchers at Washington University School of Medicine in St. Louis. They have killed the bug — methicillin-resistant Staphylococcus aureus (MRSA) — in test tubes and laboratory mice, and believe the same strategy may work in people.
Drug-resistant bacteria possess natural ability to become vulnerable to antibiotics
Infections with one of the most troublesome and least
understood antibiotic-resistant “superbugs” are increasing at alarming
rates, particularly in health-care settings. But by studying A. baumannii, a frequent cause of difficult-to-treat infections in hospitals, researchers have identified a naturally occurring process that restores its vulnerability to antibiotics.
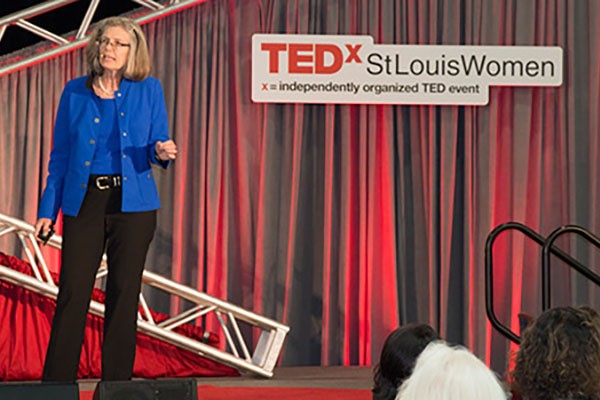
Fraser speaks about antibiotic resistance at TEDxStLouisWomen conference
Victoria J. Fraser, MD, head of the Department of Medicine, was a featured speaker at TEDxStLouisWomen, Thursday, May 28. The presentations were part of TEDWomen, a national conference focused on women and girls as creators and change-makers. Fraser spoke about antibiotic resistance and its evolution into a public health crisis.
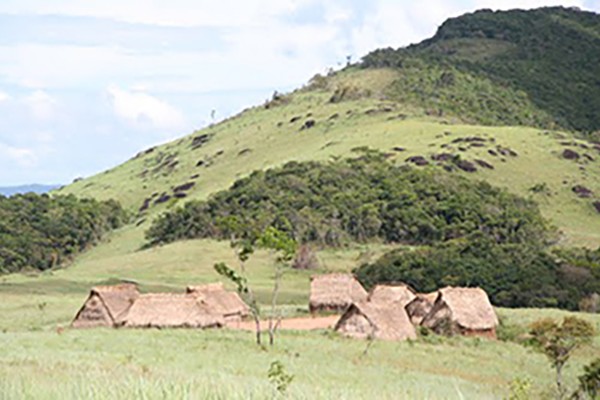
Bacterial flora of remote tribespeople carries antibiotic resistance genes
Scientists, including researchers from Washington University School of Medicine, have found antibiotic resistance genes in the bacterial flora of a South American tribe that never before had been exposed to antibiotic drugs. The findings suggest that bacteria in the human body have had the ability to resist antibiotics since long before such drugs were ever used to treat disease.
Common bacteria on verge of becoming antibiotic-resistant superbugs
Antibiotic resistance is poised to spread rapidly around the globe among bacteria frequently implicated in respiratory and urinary infections in hospital settings, according to new research at the School of Medicine.
WashU Expert: New method of finding drugs more important than new antibiotic itself
It was big news this week when Nature published the discovery of a new antibiotic, teixobactin. Teixobactin, which kills bacteria by a different pathway than other antibiotics, represented the first new class of antibiotics to be discovered in 30 years. But, says, Michael S. Kinch of Washington University in St. Louis, the drug itself may be less important than the way it was found.
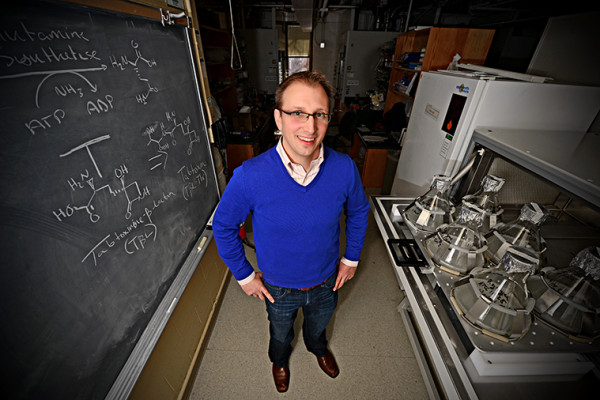
Slaying bacteria with their own weapons
A novel antibiotic delivery system would exploit small molecules called siderophores that bacteria secrete to scavenge for iron in their environments. Each bacterium has its own system of siderophores, which it pumps across its cell membrane
before releasing the iron the siderophores hold. If an antibiotic were linked to one of these scavenger molecules, it would be converted into a tiny Trojan horse that would smuggle antibiotics inside a bacterium’s cell membrane.
Soil bacteria may provide clues to curbing antibiotic resistance
Bacteria that naturally live in the soil have a vast collection of genes to fight off antibiotics, but they are much less likely to share these genes than infectious bacteria, a new study by researchers at the School of Medicine has revealed. Shown is senior author Gautam Dantas, PhD.
New drugs for bad bugs
Washington University in St Louis chemist Timothy Wencewicz says we’ll stay ahead of antibiotic resistance only if we find drugs with new scaffolds, or core chemical structures. One promising candidate, an antibiotic made by a bacterium than infects plants, caught his attention because it contains an “enchanted ring,” the beta-lactam ring that is found in penicillin. In this drug candidate, however, it acts against a different target than the penicillins.
Gut microbes in healthy kids carry antibiotic resistance genes
Friendly microbes in the intestinal tracts of healthy American children have numerous antibiotic resistance genes, according to results of a pilot study by scientists at the School of Medicine. The genes are cause for concern because they
can be shared with harmful microbes, interfering with the effectiveness of antibiotics in ways that can contribute to serious illness and, in some cases, death. Pictured is the study’s senior author, Gautum Dantas, PhD.
View More Stories