As public health officials in China brace for a potential resurgence in SARS (Sudden Acute Respiratory Syndrome) in connection with Chinese New Year on February 9, researchers at Washington University in St. Louis have published insights into a new protein that could be an important contributor to the SARS virus’ ability to cause disease and death.
When the SARS virus first jumped from its still unknown animal reservoir to humans in late 2002, it caused approximately 800 deaths and 8,000 infections in several Asian countries. Months before SARS became available for direct study, Washington University researchers used the SARS genome, finished a mere half-year after the first human infections, to look for potentially important viral proteins.
As reported recently in the journal Structure, their efforts revealed orf7a, a SARS protein that makes its home in a structure cells use to modify proteins destined for transport to the cell surface. Researchers don’t have any hard evidence of orf7a’s function, but they speculate that the protein’s positioning could enable it to block proteins that help alert the immune system or to assist reproduced viral proteins as they make their way out of the cell.
The finding exemplifies a primary directive of the University’s BioMed 21 Initiative: harness the power of genetic information to rapidly advance possibilities for patient treatment. If further investigation shows orf7a is important to the SARS virus’ ability to cause disease, scientists will already have much of the knowledge they need to develop ways to disable or weaken it.

“The key to studying novel viruses used to be the ability to isolate the virus and grow it in a culture,” notes lead author Daved Fremont, Ph.D., assistant professor of biochemistry and molecular biophysics and of pathology and immunology. “Now we can take these viral genetic sequences, which can be attained very rapidly, and get a head start on detailed biophysical investigations of emerging diseases months before we can even begin to work with such diseases in culture.”
Fremont led the initial analysis, which was dedicated to identifying sequences of genetic code that might translate into proteins that could help the virus avoid immune system attacks. He and his colleagues sought such sequences in portions of the virus’ accessory genome—areas of its genetic code not commonly found in other viruses.
SARS belongs to a class of viruses known as coronaviruses. Because those viruses typically aren’t harmful to humans, researchers suspect a gene in the SARS accessory genome may be providing the virus with most of its pathogenic punch.

Their computerized search highlighted a section of the SARS genetic code that shared several features with immune evasion proteins previously identified in other viruses. Some genetic codes contain sequences of protein-building instructions that the organisms never use, and the researchers had no proof yet that SARS actually made use of the sequence their analysis identified. Geneticists call such sequences of genetic information open reading frames, and the SARS sequence therefore came to be known as open reading frame 7a, or orf7a.
To produce the orf7a protein, research instructor Christopher Nelson, Ph.D., and graduate student Chung Lee, both of the Fremont laboratory, transplanted orf7a into bacteria. When the bacteria made the protein, researchers purified it and used it to determine the three-dimensional structure of orf7a. This purified protein also allowed colleague Michael Diamond, M.D., Ph.D., assistant professor of molecular microbiology, to generate orf7a-specific antibodies.
Andrew Pekosz, Ph.D., assistant professor of molecular microbiology, applied those antibodies to cell cultures infected with the SARS virus, revealing that the virus does produce the orf7a protein.

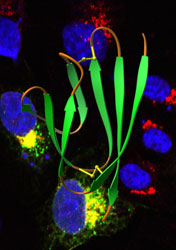
“We learned that the virus does make the protein, but we also got a big surprise when we looked at where the protein was,” says Diamond. “Based on our analysis of the genome, we thought the orf7a protein would be on the surface of infected cells, decoying the immune system in some obvious way. But we found that most orf7a protein was inside the cell and little if any of it was on the surface.”
Further experiments revealed that part of the protein specializes in getting orf7a to the Golgi apparatus—the structure cells use to prepare proteins for transport to the cell surface–and keeping it there.
Plans for follow-up include studies of a recently produced version of the SARS virus where orf7a has been genetically knocked out. Scientists hope to use it to learn what orf7a does for the virus and how important it is to the virus’ ability to survive and reproduce.
“If knocking out orf7a significantly attenuates the virus, we may able to look at the possibility of using it to make a SARS vaccine,” Pekosz speculates.
Nelson CA, Pekosz A, Lee CA, Diamond MS, Fremont DH. Structure and intracellular targeting of the SARS-Coronavirus Orf7a Accessory Protein. Structure, January 2005, 75-85.
Funding from the National Institutes of Health supported this research.
Washington University School of Medicine’s full-time and volunteer faculty physicians also are the medical staff of Barnes-Jewish and St. Louis Children’s hospitals. The School of Medicine is one of the leading medical research, teaching and patient care institutions in the nation, currently ranked second in the nation by U.S. News & World Report. Through its affiliations with Barnes-Jewish and St. Louis Children’s hospitals, the School of Medicine is linked to BJC HealthCare.