Human diseases and social networks seem to have little in common. However, at the crux of these two lies a network, communities within the network, and farther even, substructures of the communities. In a recent paper in Physical Review E 77:016104 (2008), Weixiong Zhang, Ph.D., Washington University associate professor of computer science and engineering and of genetics, along with his Ph.D. student, Jianhua Ruan, published an algorithm (a recipe of computer instructions) to automatically identify communities and their subtle structures in various networks.
Many complex systems can be represented as networks, Zhang said, including the genetic networks he studies, social networks and the Internet itself. The community structure of networks features a natural division in which the vertices in each subnetwork are highly involved with each other, though connected less strongly with the rest of the network. Communities are relatively independent of one another structurally, but researchers think that each community may correspond to a fundamental functional unit. A community in a genetic network usually contains genes with similar functions, just as a community on the World Wide Web often corresponds to Web pages on similar topics.
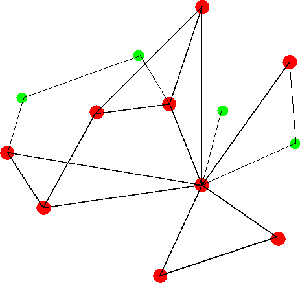
All Zhang and Ruan need are data. Their algorithm is more scalable than existing similar algorithms and can detect communities at a finer scale and with a higher accuracy. One impact of having such a computational biology tool is found in the genomics field. Using this tool, researchers may be better able to identify and understand communities of genes and their networks as well as how they cooperate in causing diseases, such as sepsis, virus infections, cancer and Alzheimer’s disease.
Versatile math tool
Zhang and Ruan’s algorithm is so versatile that it has been applied to identify the community structure of a network of co-expressed genes involved in bacterial sepsis.
“This is a tool not only for biological research, but also for sociological research,” Zhang said. It can determine, for instance, how people interact in social networks and how scientists collaborate in scientific research.
In biological systems there are lots of communities with many proteins involved to form complexes. “We can use this tool to identify structures embedded in the data,” Zhang said. “We’ve identified the substructures of three different RNA polymerase complexes from noisy data, for instance, which are crucial for gene transcription.”
Zhang began his computer science career as a specialist in artificial intelligence, but in recent years he has focused more on computational biology. His goal is to use computational means to solve some basic biology problems and those related to human diseases. For example, his group studied a basic problem of the transcription mechanism of microRNAs, which are small, noncoding RNAs that regulate the development and stress responses of nearly all eukaryotic species that have been studied. Using machine learning techniques, Zhang and his collaborators showed that almost all intergenic microRNA genes in four model species, human, mouse, rice and mustard plant (Arabidopsis), are transcribed by RNA polymerase II, which transcribes protein-coding genes. The results were published in PLoS Computational Biology, 3(3):e37 (2007).
Multidisciplinary research that combines computational approaches with biological data is a hallmark of research themes in Zhang’s group. As another example, in a paper published in Genome Biology, 7(6):R49 (2006), Zhang and his Ph.D. student, Guandong Wang, developed an algorithm called WordSpy that identifies cis-regulatory elements — short DNA sequences that are critical to the regulation of gene expression — from a large amount of genome sequences.
Stealth from the ancient Greeks
WordSpy was inspired by an old information-hiding technique called stegography, which can be traced back to ancient Greece. As such, their method can be used to analyze not only genomic sequences, but also natural languages. In fact, their method has been extended to segment words and phrases in Chinese.
Aside from studying networks, Zhang also has formed a broad network of collaborations with scientists across the WUSTL campus and outside of the university. The problems he studies are diverse, ranging from stress responses and virus infection in plants, such as rice, to human diseases, including Alzheimer’s disease, herpes virus infection, sepsis, cardiac hypertrophy, lung cancer and lung transplantation. The computational tools his group has developed are helping him and his collaborators come to grips with how perturbation to gene expression can lead to complex traits and human diseases as well as how microRNAs regulate gene expression.
Zhang recently was awarded a grant from the Alzheimer’s Association to develop computational systems biology methods for analyzing gene expression perturbation in diseased brains. He has been collaborating with scientists in the Washington University School of Medicine and Scripps Institute in La Jolla, Calif., to study roughly 30 postmortem brain samples of people who died from Alzheimer’s disease.
“I’m interested in modeling gene expression perturbation in diseased brains and am looking for the genetic signature,” Zhang said. “Due to the complexity of Alzheimer’s disease, we are developing other tools. It’s a polygenic disease, with a lot of genes at work. I’m sure we’ll find that a network is involved.”