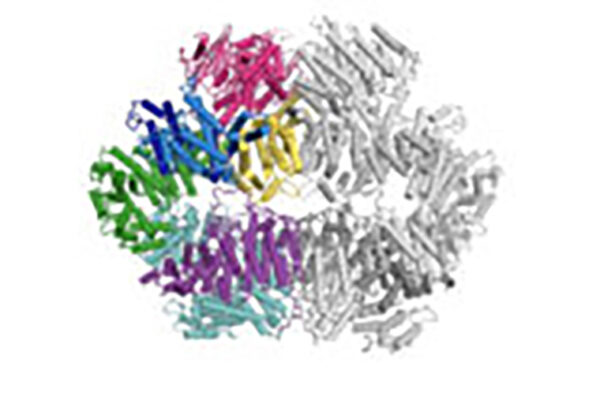
Shown is the first look at the Folding@home project’s simulations of the COVID-19 spike protein. The three colors represent components of the spike protein; this is the protein that the novel coronavirus uses to infect cells. The site where the protein binds to human cells, to infect them, is on the top of the protein. Using Folding@home, the researchers are aiming to develop an accurate picture of what happens during infection. Understanding these details could help reveal ways to block the virus from infecting cells. (Video: Greg Bowman and Maxwell Zimmerman)
People around the world are isolating themselves to help slow the spread of COVID-19. But there is another way those confined to their homes — but connected online — can join the fight against the novel coronavirus. Among the research programs racing to develop therapies and vaccines for this new pandemic virus is one of the largest crowdsourced supercomputing projects in the world.
Led by computational biophysicist Greg Bowman, associate professor of biochemistry and molecular biophysics at Washington University School of Medicine in St. Louis, the project is called Folding@home. It relies on the collective power of volunteers’ home computers to perform the complex calculations required to simulate protein dynamics.
Volunteers from all over the world can install a software program that runs those calculations when a computer otherwise would sit idle. Often motivated by personal experience with various diseases, the participants get to select an area of contribution, such as boosting cancer research, preventing Alzheimer’s disease or — now — fighting the novel coronavirus.
For example, Bowman and his team are trying to understand the structure of COVID-19’s spike protein, which is what the virus uses to infect cells. Such research could reveal ways to block the protein and, consequently, infection. Since announcing in late February the project’s new focus on coronavirus, the number of Folding@home volunteers has skyrocketed, with some 400,000 new folders joining the effort, Bowman said.
“The response so far has been overwhelming and wonderful, but there is always more useful science to be done,” he said. “Understanding all the various shapes that the spike protein can take on as its molecules bounce and shift can lead to the development of new drugs that can block it, stopping the virus from infecting more cells. We are continuing to scale up our research as fast as we can.”
Bowman has shared what could be thought of as a first glimpse of the moving COVID-19 spike protein. It consists of three different proteins that fit together like a 3D puzzle. The simulation reveals a pocket that helps the virus bind to human cells and infect them.
In the spirit of open science and the rapid sharing of new knowledge about COVID-19, Bowman said the research team will publish findings on free and open-access preprint sites, such as bioRxiv.
To join the effort and put your computer to work against coronavirus, visit https://foldingathome.org.
Washington University School of Medicine’s 1,500 faculty physicians also are the medical staff of Barnes-Jewish and St. Louis Children’s hospitals. The School of Medicine is a leader in medical research, teaching and patient care, ranking among the top 10 medical schools in the nation by U.S. News & World Report. Through its affiliations with Barnes-Jewish and St. Louis Children’s hospitals, the School of Medicine is linked to BJC HealthCare.
WashU Response to COVID-19
Visit coronavirus.wustl.edu for the latest information about WashU updates and policies. See all stories related to COVID-19.