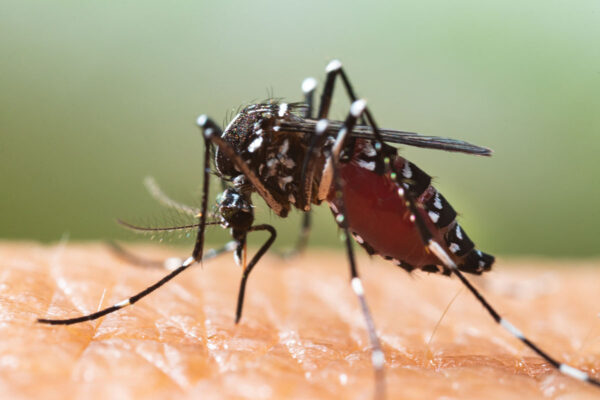
Chikungunya virus, once confined to the Eastern Hemisphere, has infected more than 1 million people in the Americas since 2013, when mosquitoes carrying the virus were discovered in the Caribbean. Most people who become infected develop fever and joint pain that last about a week. But in up to half of patients, the virus can cause severe arthritis that persists for months or years. There is no treatment to prevent the short-lived infection from persisting into chronic arthritis.
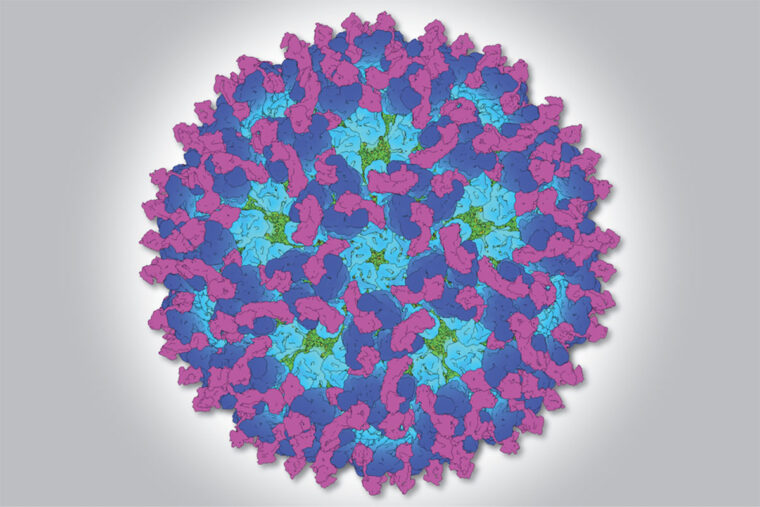
Now, researchers have uncovered information that could help stop the debilitating condition. A team at Washington University School of Medicine in St. Louis has snapped high-resolution pictures of the virus latched onto a protein found on the surface of cells in the joints. The protein used in the study was taken from mice, but people have the same protein, and the virus interacts with the mouse and human proteins in virtually identical ways.
The structures, published May 9 in the journal Cell, shows in atomic-level detail how the virus and cell-surface protein fit together – data that promises to accelerate efforts to design drugs and vaccines to prevent or treat arthritis caused by chikungunya or related viruses.
“Chikungunya arthritis comes on very suddenly and can be very painful – people can barely walk around – and we have nothing specific to treat or prevent it,” said co-senior author Michael S. Diamond, MD, PhD, the Herbert S. Gasser Professor of Medicine, and a professor of molecular microbiology, and of pathology and immunology.
“Now that we have these new structures, we can see how to disrupt the interaction between the virus and the protein it uses to get inside cells in the joints and other musculoskeletal tissues in order to block infections,” added co-senior author Daved Fremont, professor of pathology and immunology, of biochemistry and molecular biophysics, and of molecular microbiology.
Chikungunya and its cousins – Mayaro, Ross River and O’nyong-nyong viruses – belong to a family of alphaviruses that are spread by mosquitoes and cause joint pain. In recent years, such viruses have been infecting people and animals in ever larger regions of the globe. In 2018, Diamond, Fremont and colleagues including postdoctoral researcher Rong Zhang identified the protein Mxra8, which is found on the outer surface of cells in the joints, as the molecular handle that chikungunya and related viruses grasp to gain entry into cells of mice, humans and other species. The human and mice versions of Mxra8 are 79 percent identical, and chikungunya virus interacts with both versions in the same way. Viruses need to latch onto the protein to cause disease; in mice, thwarting chikungunya’s attempts to attach to the protein using blocking antibodies or decoy receptors reduced signs of arthritis.
To design effective drugs and vaccines that interfere with attachment, researchers need a detailed picture of the molecular interactions between the virus and the protein. Fremont worked with graduate students Katherine Basore, the paper’s first author, and Arthur Kim, to visualize the virus bound to the cell-surface protein. The researchers used a technique called cryo-electron microscopy (cryo-EM) at Washington University’s Center for Cellular Imaging, which installed a state-of-the-art cryo-EM machine in 2018.
Images were obtained using a chikungunya virus-like particle – which has the shape of a virus but cannot cause infections because it carries no genetic material inside – as well as fully infectious chikungunya virus. The virus-like particles are being evaluated in clinical trials as a potential vaccine for chikungunya.
To visualize how the virus interacts with the cell-surface protein, the researchers first flash-froze the viral particles attached to the protein. The snap freeze was necessary to keep the particles from being destroyed during the experiment. Then, the researchers shot a beam of electrons through the sample, mapped where the electrons landed on a detector, and used computer programs to reconstruct the electron density patterns and thereby the 3D structure of the viral particles bound to the cell-surface protein.
“Our cryo-EM maps allow us to view the whole virus particle with Mxra8 bound to it, but not at a high enough resolution to pinpoint precise atomic locations,” Basore said. “So we used existing high-resolution X-ray crystal structures of the components of the virus, in addition to our own crystal structure of Mxra8, to build an atomic model of the entire assembly. This allowed us to see the full scope of these interactions that we couldn’t achieve from either X-ray crystallography or cryo-EM alone.”
The high-resolution structure will aid efforts to screen experimental drugs for their ability to block attachment to the protein on cells in the joints, evaluate whether the antibodies elicited by investigational vaccines are likely to prevent infection, and analyze whether mutations in viruses affect their virulence.

Comments and respectful dialogue are encouraged, but content will be moderated. Please, no personal attacks, obscenity or profanity, selling of commercial products, or endorsements of political candidates or positions. We reserve the right to remove any inappropriate comments. We also cannot address individual medical concerns or provide medical advice in this forum.